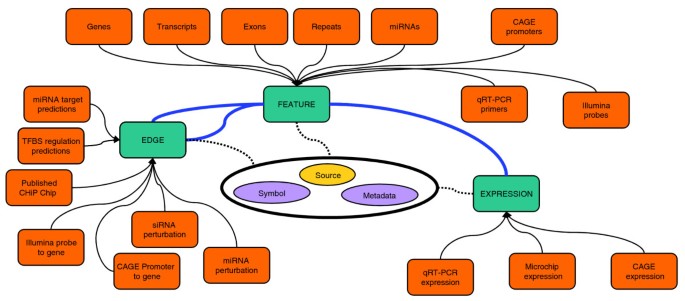
FANTOM4 EdgeExpressDB: an integrated database of promoters, genes, microRNAs, expression dynamics and regulatory interactions | Genome Biology | Full Text
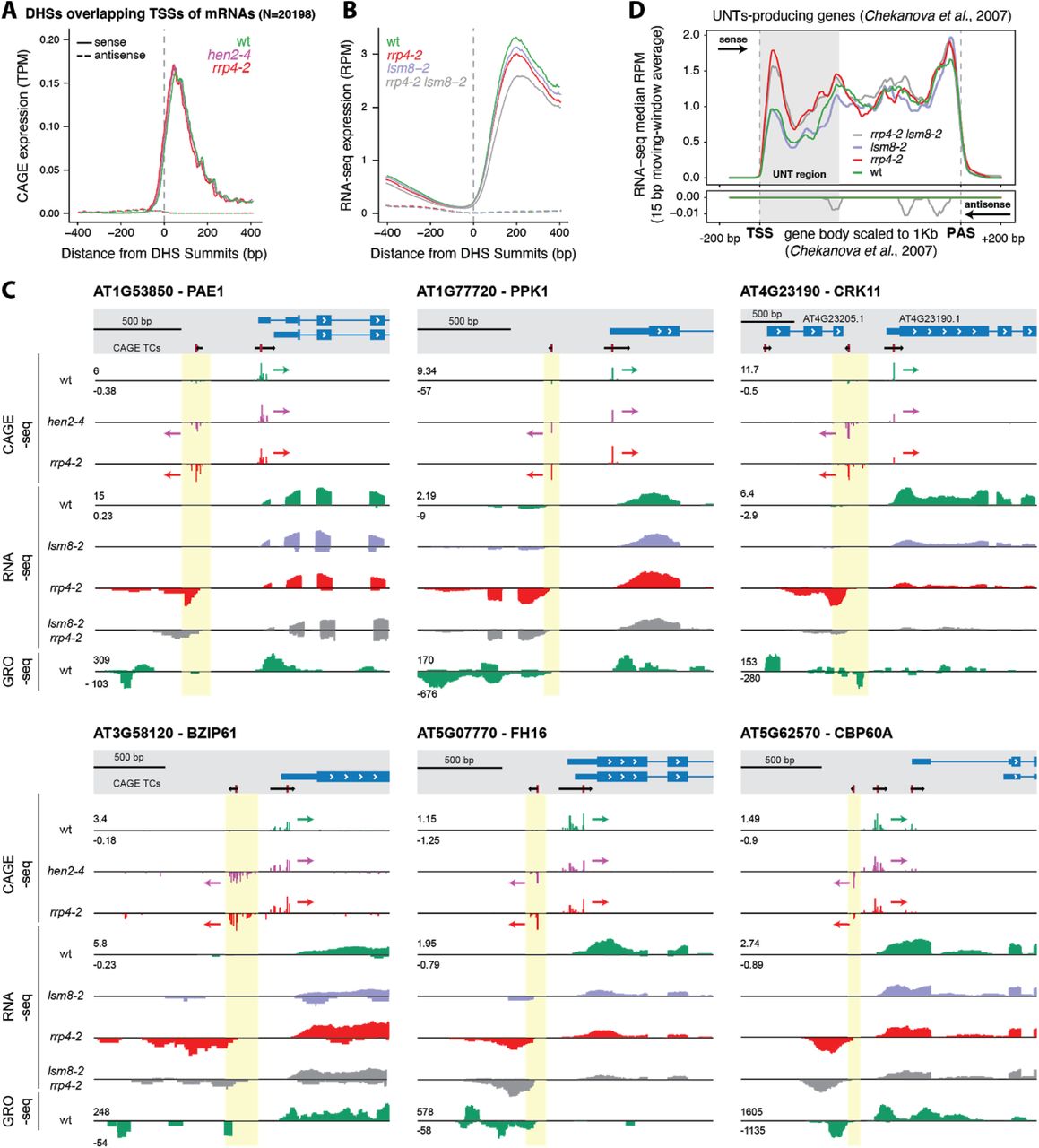
Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv
Distal CpG islands can serve as alternative promoters to transcribe genes with silenced proximal promoters

Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv
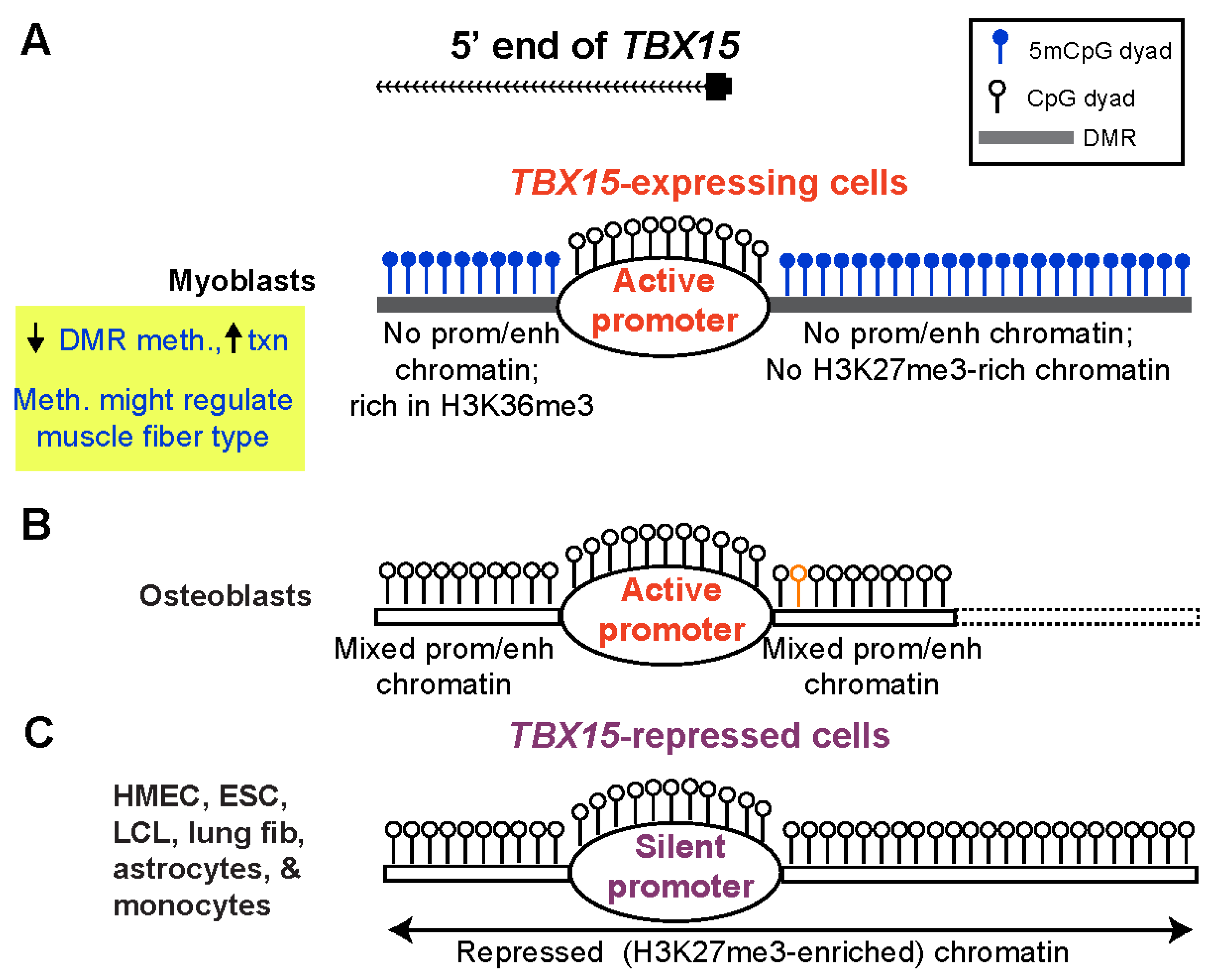
Epigenomes | Free Full-Text | Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage

The miR-430 locus with extreme promoter density forms a transcription body during the minor wave of zygotic genome activation - ScienceDirect
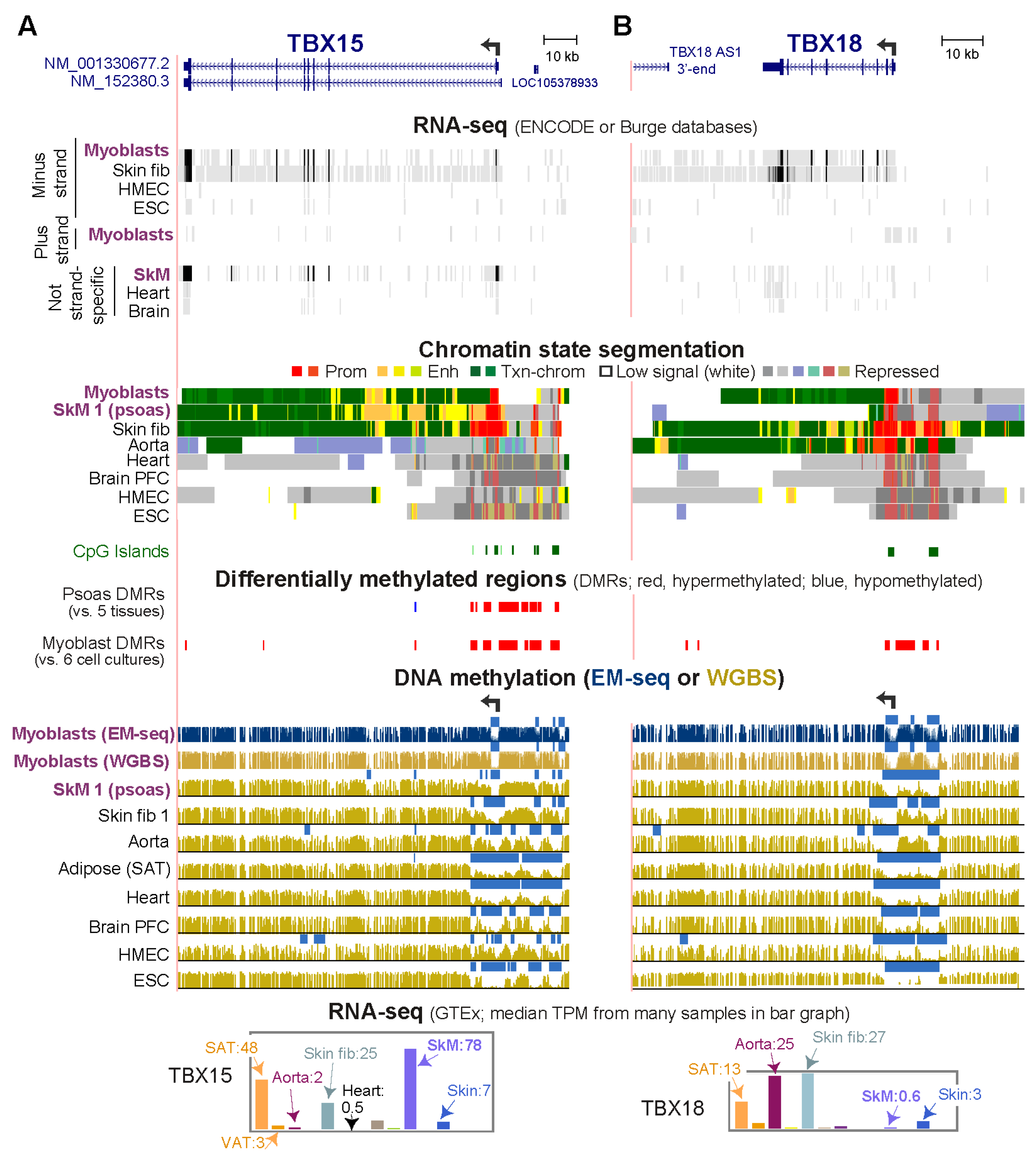
Epigenomes | Free Full-Text | Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for analysis of CAGE data | bioRxiv

Multiple transcription initiation and alternative promoter usage in G.... | Download Scientific Diagram

Cap analysis of gene expression reveals alternative promoter usage in a rat model of hypertension | Life Science Alliance
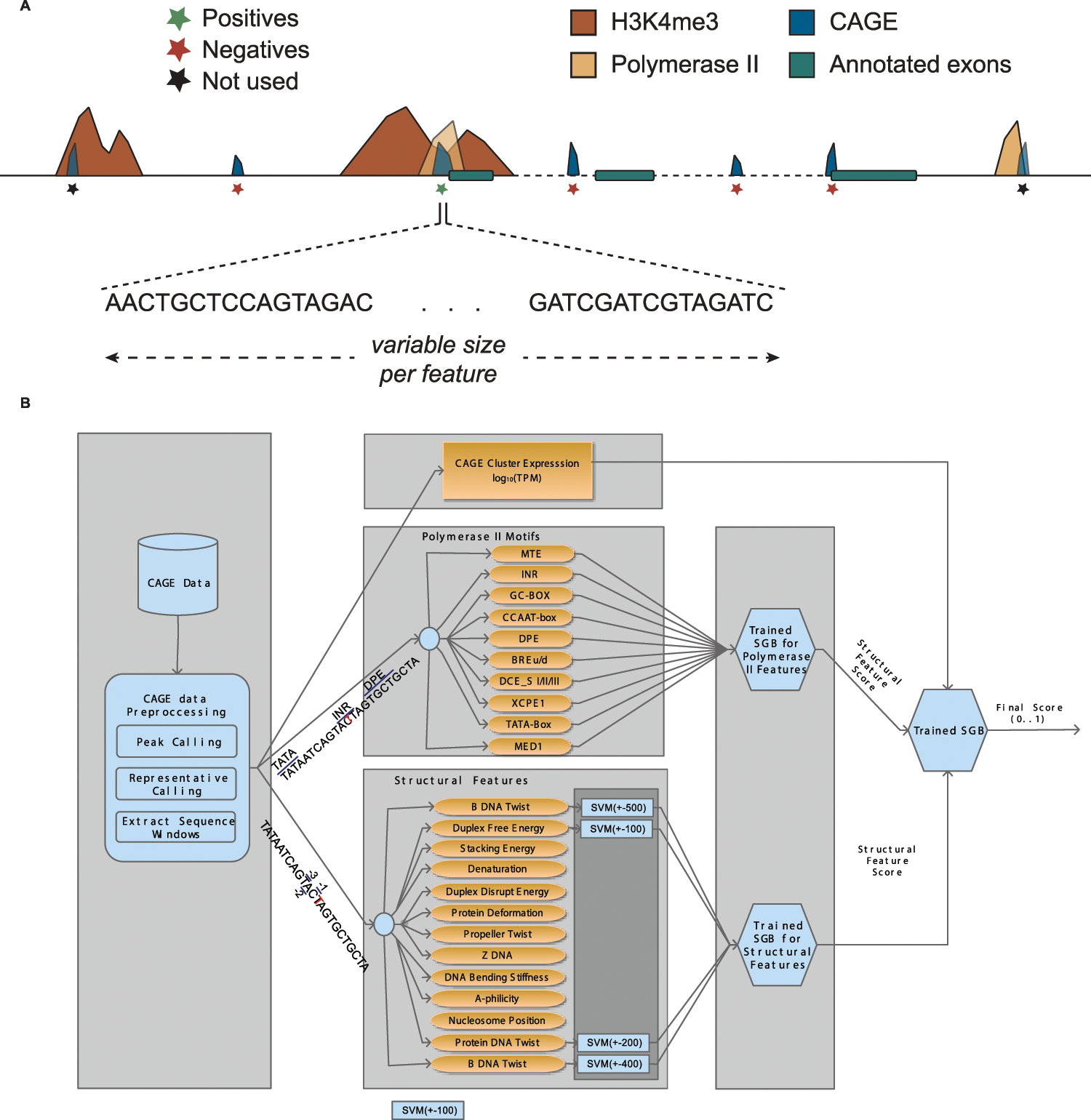
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports